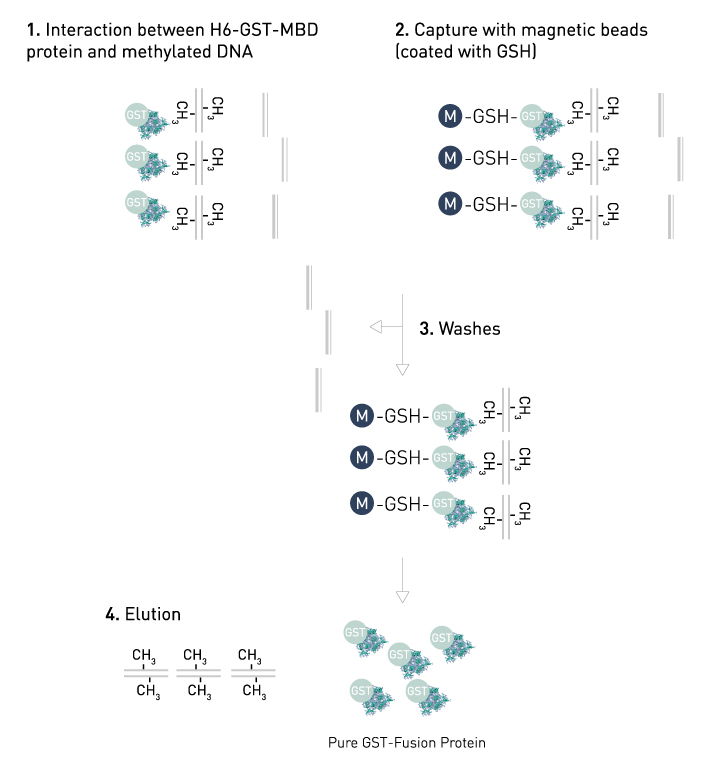
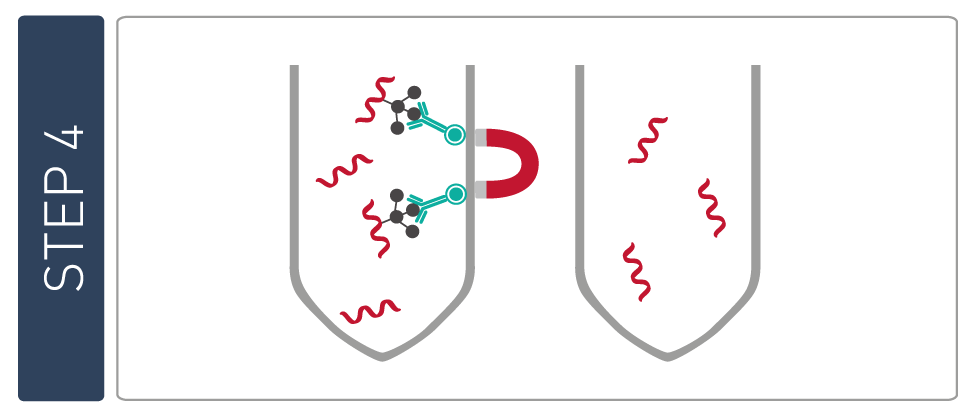
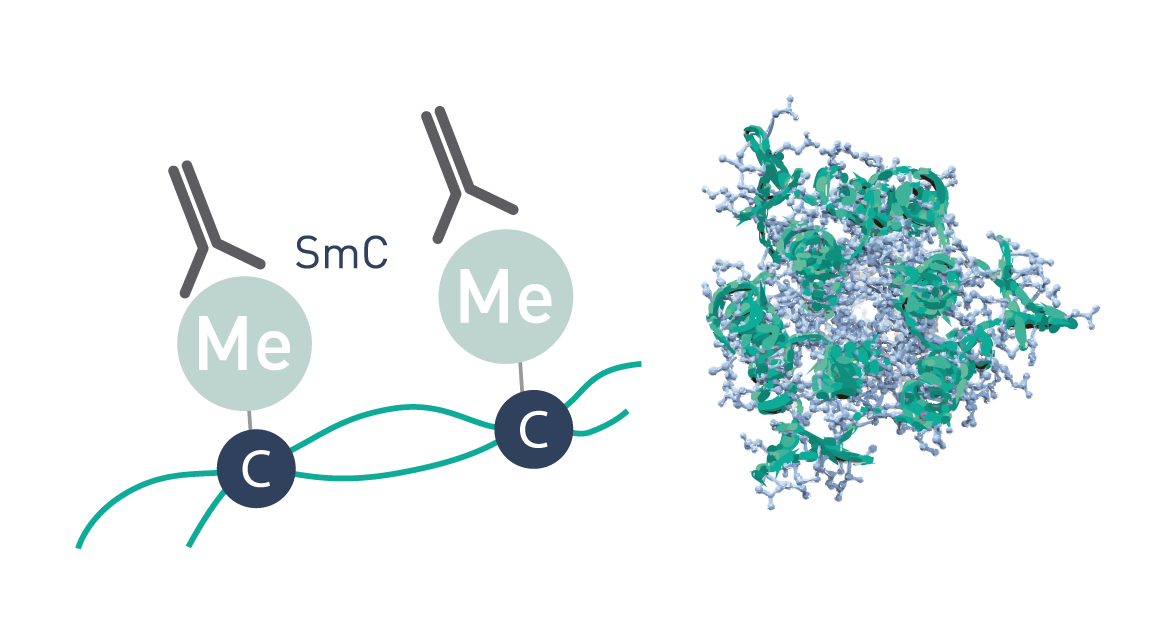
The MBD technology used in our MethylCap Kit is based on the very high affinity of a H6-GST-MBD fusion protein for methylated DNA. This protein consists of the methyl binding domain (MBD) of human MeCP2, as a C-terminal fusion with Glutathione-S-transferase (GST) containing an N-terminal His6-tag. The H6-GST-MBD fusion protein can be used to specifically isolate DNA containing methylated CpGs.
Diagenode’s MethylCap Kit enables high enrichment of double-stranded DNA and a differential fractionation in function of the methylated CpG density. Fractionation reduces the complexity of samples and makes subsequent next generation sequencing easier. Prior to the MethylCap assay, DNA is first extracted and sheared using the Bioruptor® Sonicator.
ADVANTAGES
- Unaffected DNA
- Robust & reproducible technique
- NGS compatible