Histones are the main constituents of the protein part of chromosomes of eukaryotic cells. They are rich in the amino acids arginine and lysine and have been greatly conserved during evolution. Histones pack the DNA into tight masses of chromatin. Two core histones of each class H2A, H2B, H3 and H4 assemble and are wrapped by 146 base pairs of DNA to form one octameric nucleosome. Histone tails undergo numerous post-translational modifications, which either directly or indirectly alter chromatin structure to facilitate transcriptional activation or repression or other nuclear processes. In addition to the genetic code, combinations of the different histone modifications reveal the so-called “histone code”. Histone methylation and demethylation is dynamically regulated by respectively histone methyl transferases and histone demethylases.
H3K4me1 Antibody
(pAb-037-050)
Get a quote
Request a quote for a bulk order for H3K4me1 Antibody. Please fill out the form here below. Your local sales account manager will get in touch with you shortly and send you a quotation based on your requirements.
Bulk order
| Lot | A1657D |
|---|---|
| Concentration | 2.9 µg/µl |
| Species reactivity | Human, mouse, pig |
| Type | Polyclonal. ChIP-grade. ChIP-seq grade. CUT&Tag-grade. |
| Purity | Affinity purified polyclonal antibody. |
| Host | Rabbit |
| Storage Conditions | Store at -20°C; for long storage, store at -80°C. Avoid multiple freeze-thaw cycles. |
| Storage Buffer | PBS containing 0.05% azide and 0.05% ProClin 300. |
| Precautions | This product is for research use only. Not for use in diagnostic or therapeutic procedures. |
| Applications | Suggested dilution | References |
|---|---|---|
| ChIP/ChIP-seq * | 1-2 μg/IP | Fig 1, 2 |
| CUT&Tag | 1μg | Fig 3 |
| ELISA | 1:500 | Fig 4 |
| Dot Blotting | 1:10,000 | Fig 5 |
| Western Blotting | 1:500 | Fig 6 |
| Immunofluorescence | 1:500 | Fig 7 |
* Please note that the optimal antibody amount per IP should be determined by the end-user. We recommend testing 1-5 μg per IP.
- Validation Data
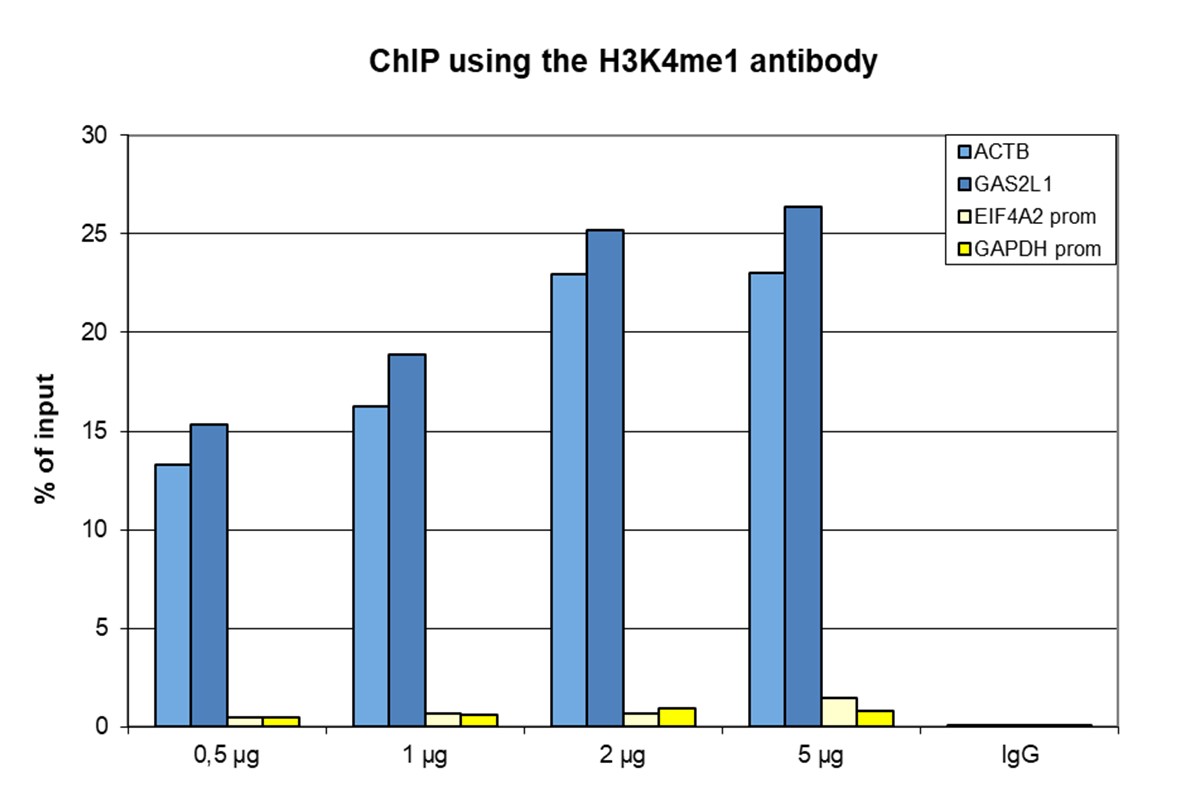
Figure 1. ChIP results obtained with the antibody directed against H3K4me1
ChIP was performed with the antibody against H3K4me1 (cat. No. C15410037) on sheared chromatin from 500,000 HeLaS3 cells using the iDeal ChIP-seq kit (cat. No. C01010051). A titration of the antibody consisting of 0.5, 1, 2 and 5 µg per ChIP experiment was analysed. IgG (2 µg/IP) was used as negative IP control. Quantitative PCR was performed with primers for a region surrounding the ACTB and GAS2L1 genes, used as positive controls, and for the promoters of the GAPDH and EIF4A2 genes, used as negative controls. Figure 1 shows the recovery, expressed as a % of input (the relative amount of immunoprecipitated DNA compared to input DNA after qPCR analysis).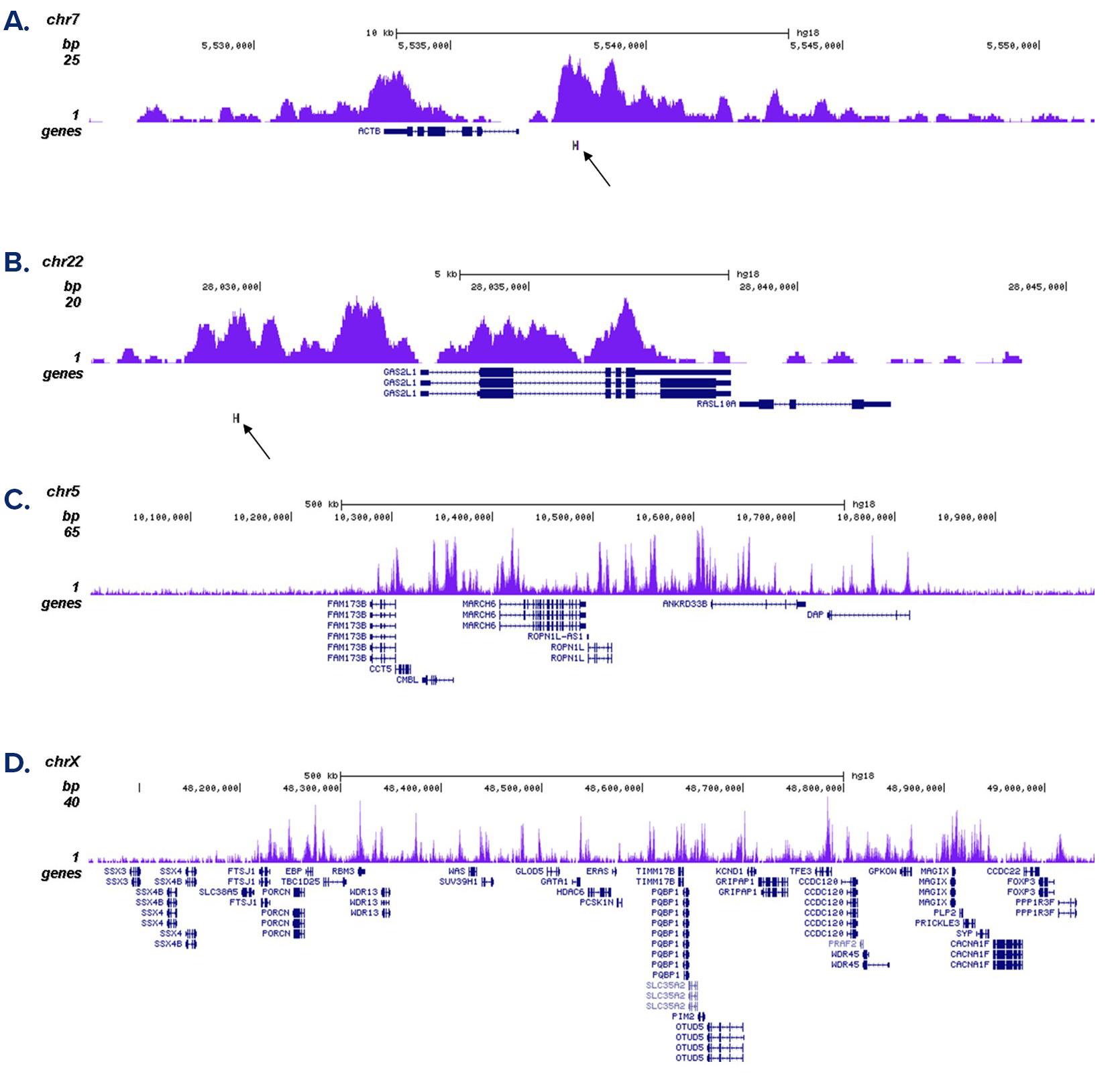
Figure 2. ChIP-seq results obtained with the antibody directed against H3K4me1
ChIP was performed as described above with 1 µg of the antibody against H3K4me1 (cat. No. C15410037). The IP’d DNA was subsequently analysed on an Illumina Genome Analyzer. Library preparation, cluster generation and sequencing were performed according to the manufacturer’s instructions. The 36 bp tags were aligned to the human genome using the ELAND algorithm. Figure 2A and B show the enrichment in chromosomal regions surrounding the ACTB and GAS2L1 positive control genes. The position of the amplicon used in the qPCR validation is indicated by an arrow. Figure 2C and D show the H3K4me1 signal in two 1 Mb regions of chromosome 5 and X, respectively.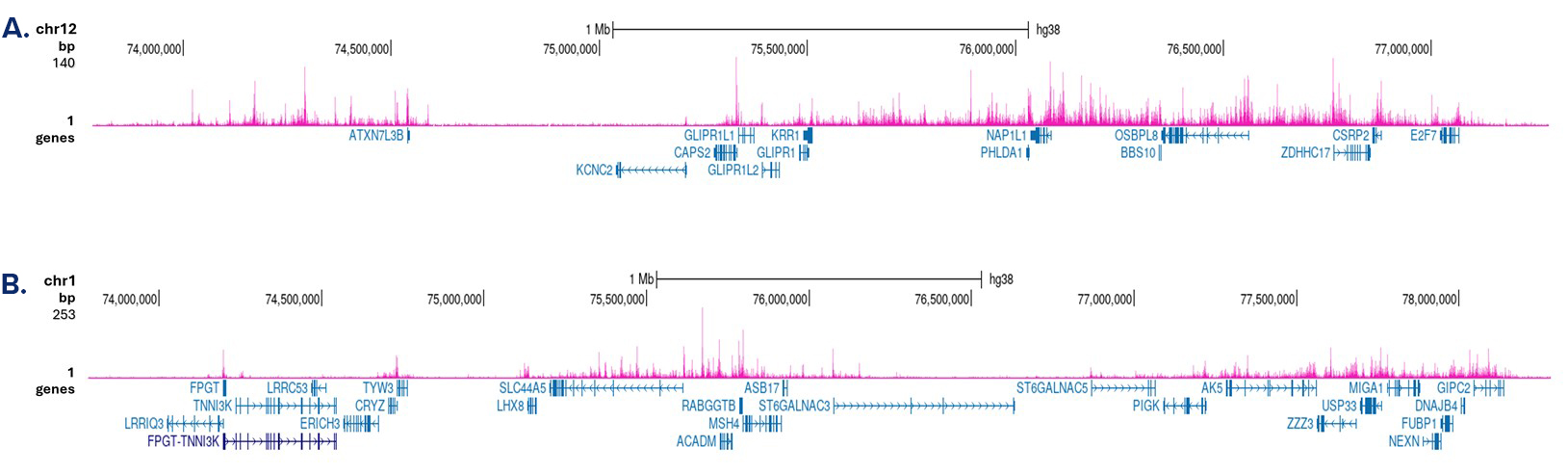
Figure 3. Cut&Tag results obtained with the antibody directed against H3K4me1
Cut&Tag was performed on 50,000 K562 cells using 1 µg of the antibody against H3K4me1 (cat. No. C15410037) and the iDeal CUT&Tag kit (cat. No. C01070020). The libraries were subsequently analysed on an Illumina NovaSeq sequencer (2x50 bp paired-end reads) according to the manufacturer’s instructions. The tags were aligned to the human genome (hg38) using the BWA algorithm. Figure 3 shows the peak distribution in two genomic regions on chromosome 12 and 1 (figure 3A and B, respectively).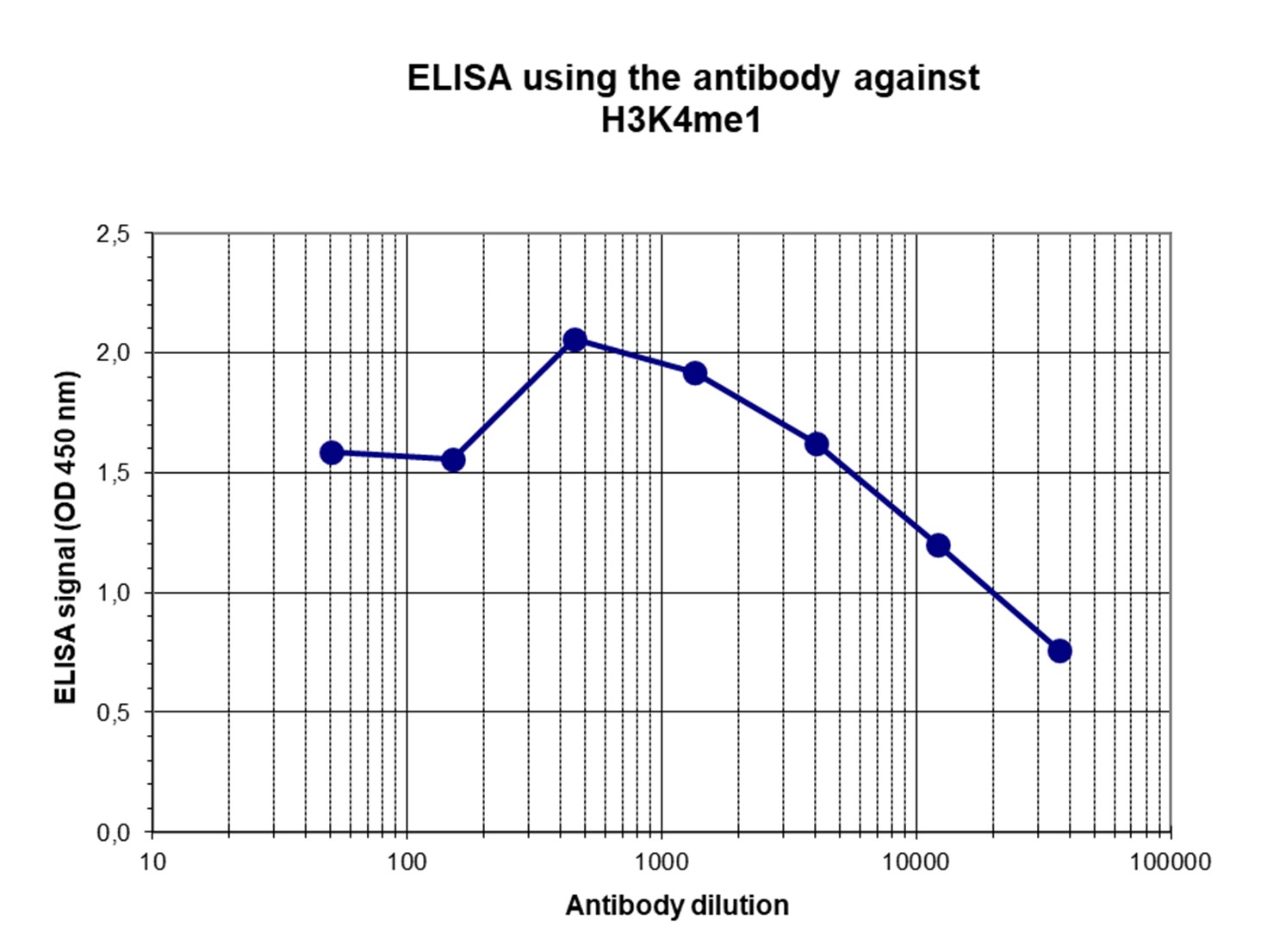
Figure 4. Determination of the titer
To determine the titer, an ELISA was performed using a serial dilution of the antibody directed against H3K4me1 (cat. No. C15410037) in antigen coated wells. The antigen used was a peptide containing the histone modification of interest. By plotting the absorbance against the antibody dilution (Figure 4), the titer of the antibody was estimated to be 1:20,100.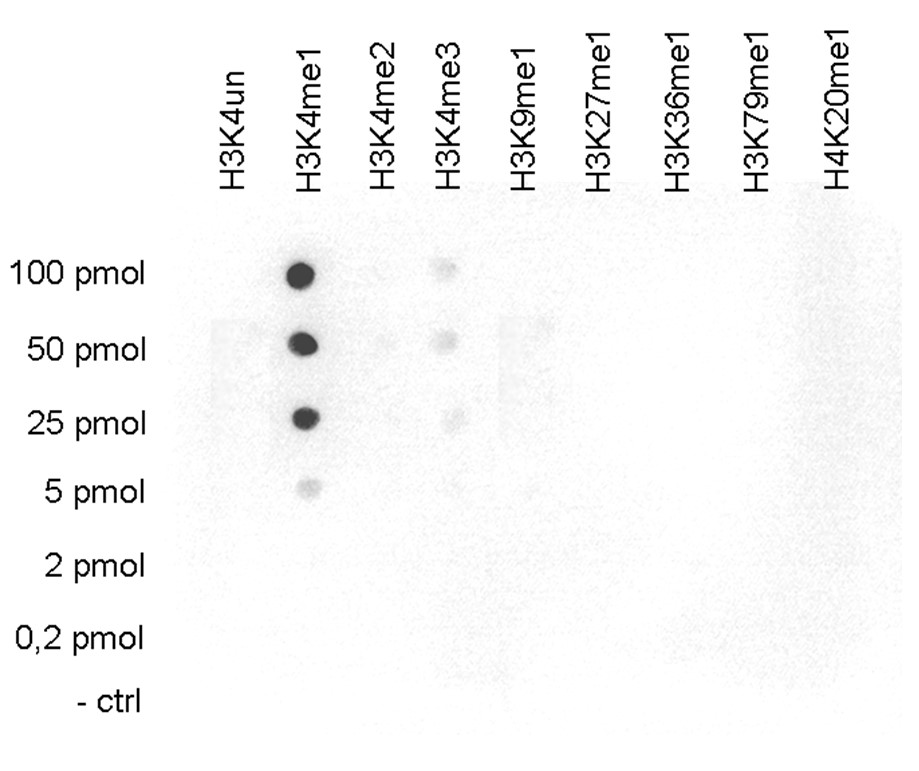
Figure 5.Cross reactivity tests using the antibody directed against H3K4me1
A Dot Blot analysis was performed to test the cross reactivity of the antibody against H3K4me1 (cat. No. C15410037) with peptides containing other modifications or unmodified sequences of histone H3. One hundred to 0.2 pmol of the respective peptides were spotted on a membrane. The antibody was used at a dilution of 1:10,000. Figure 5 shows a high specificity of the antibody for the modification of interest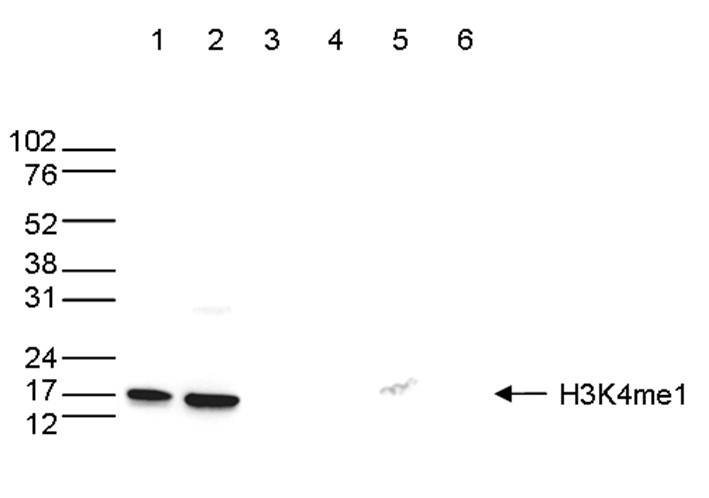
Figure 6. Western blot analysis using the antibody directed against H3K4me1
Western blot was performed on whole cell (25 µg, lane 1) and histone extracts (15 µg, lane 2) from HeLa cells, and on 1 µg of recombinant histone H2A, H2B, H3 and H4 (lane 3, 4, 5 and 6, respectively) using the antibody against H3K4me1 (cat. No. C15410037). The antibody was diluted 1:500 in TBS-Tween containing 5% skimmed milk. The position of the protein of interest is shown on the right, the marker (in kDa) is shown on the left.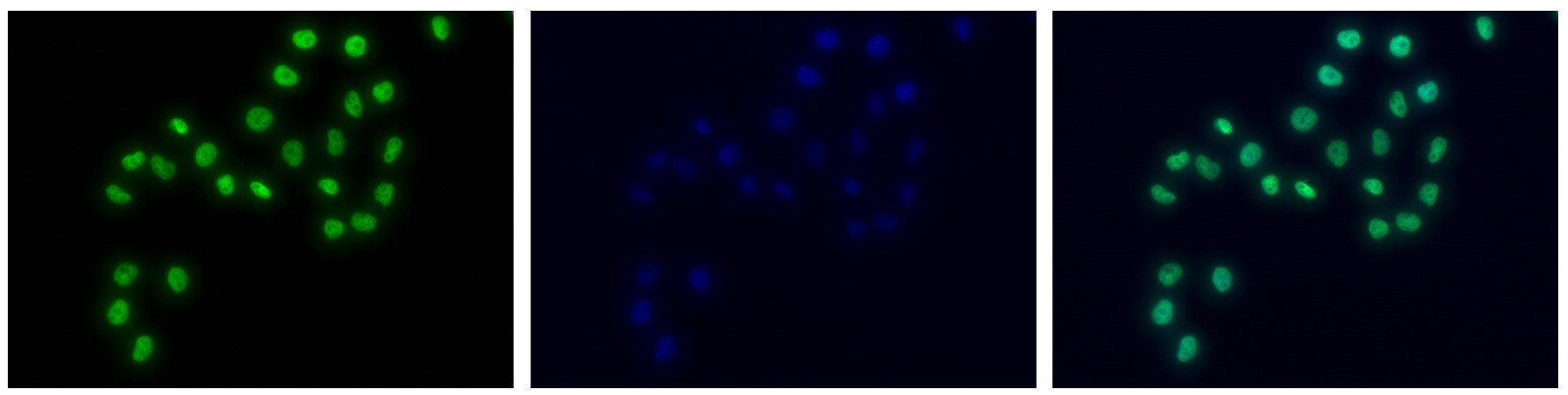
Figure 7. Immunofluorescence using the antibody directed against H3K4me1
HeLa cells were stained with the antibody against H3K4me1 (Cat. C15410037) and with DAPI. Cells were fixed with 4% formaldehyde for 10’ and blocked with PBS/TX-100 containing 5% normal goat serum and 1% BSA. The cells were immunofluorescently labeled with the H3K4me1 antibody (left) diluted 1:500 in blocking solution followed by an anti-rabbit antibody conjugated to Alexa488. The middle panel shows staining of the nuclei with DAPI. A merge of the two stainings is shown on the right. - Target Description
- Applications
ELISA
Enzyme-linked immunosorbent assay. Read moreDB
Dot blotting Read moreWB
Western blot : The quality of antibodies used in this technique is crucial for correct and specific protein identification. Diagenode offers huge selection of highly sensitive and specific western blot-validated antibodies. Learn more about: Load... Read moreChIP-seq (ab)
Read moreChIP-qPCR (ab)
Read moreCUT&Tag
The quality of antibody used in CUT&Tag is one of the crucial factors for assay success. The antibodies with confirmed high specificity will target only the protein of interest, enabling real results. Check out our selection of antibodies vali... Read more - Documents
Epigenetic Antibodies Brochure BROCHURE
More than in any other immuoprecipitation assays, quality antibodies are critical tools in many e...Download Antibodies you can trust POSTER
Epigenetic research tools have evolved over time from endpoint PCR to qPCR to the analyses of lar...Download Datasheet H3K4me1 C15410037 DATASHEET Download - Safety sheets
SDS C15410037 H3K4me1 Antibody US en Download SDS C15410037 H3K4me1 Antibody GB en Download SDS C15410037 H3K4me1 Antibody DE de Download SDS C15410037 H3K4me1 Antibody BE nl Download SDS C15410037 H3K4me1 Antibody JP ja Download SDS C15410037 H3K4me1 Antibody FR fr Download SDS C15410037 H3K4me1 Antibody ES es Download SDS C15410037 H3K4me1 Antibody BE fr Download - Publications

How to properly cite our product/service in your work
We strongly recommend using this: H3K4me1 Antibody (Hologic Diagenode Cat# C15410037 Lot# A1657D). Click here to copy to clipboard.
Using our products or services in your publication? Let us know!
Multimodal epigenetic and enhancer network remodeling shape the transcriptional landscape of human beige adipocytes
Hazell Pickering, Sarah et al.
Epigenetic regulation is a key determinant of adipocyte fate, driving the differentiation toward white or thermogenic beige phenotypes in response to environmental cues. To dissect the mechanisms orchestrating this plasticity in human adipocytes, we conducted an integrative analysis of transcriptomic, epigenomic...Multimodal epigenetic and enhancer network remodeling shape the transcriptional landscape of human beige adipocytes
Hazell Pickering, Sarah et al.
Abstract Epigenetic regulation is a key determinant of adipocyte fate, driving the differentiation toward white or thermogenic beige phenotypes in response to environmental cues. To dissect the mechanisms orchestrating this plasticity in human adipocytes, we conducted an integrative analysis of transcriptomic, epig...Extrusion fountains are restricted by WAPL-dependent cohesin release and CTCF barriers
Liu, Ning Qing et al.
Interphase chromosomes are mainly shaped by loop extrusion and compartmentalisation mechanisms. However, their temporal component and cause-effect relationships remain largely unknown. In this study, we use acute degradation of WAPL, CTCF and cohesin in mouse embryonic stem cells to investigate the dynamics of l...Single-cell multi-omics, spatial transcriptomics and systematic perturbation decode circuitry of neural crest fate decisions
Hu Z. et al.
Cranial neural crest (NC) cells, which can migrate, adopt multiple fates, and form most of the craniofacial skeleton, are an excellent model for studying cell fate decisions. Using time-resolved single-cell multi-omics, spatial transcriptomics, and systematic Perturb-seq, we fully deciphered zebrafish cranial NC pro...KRAS promotes GLI2-dependent transcription during pancreatic carcinogenesis
Sigafoos A.N. et al.
Aberrant activation of GLI transcription factors has been implicated in the pathogenesis of different tumor types including pancreatic ductal adenocarcinoma (PDAC). However, the mechanistic link with established drivers of this disease remains in part elusive. Here, using a new genetically-engineered mouse model ove...Chromatin profiling reveals TFAP4 as a critical transcriptional regulator of bovine satellite cell differentiation
Pengcheng Lyu et al.
Background Satellite cells are myogenic precursor cells in adult skeletal muscle and play a crucial role in skeletal muscle regeneration, maintenance, and growth. Like embryonic myoblasts, satellite cells have the ability to proliferate, differentiate, and fuse to form multinucleated myofibers. In this study, we ai...RUNX1 colludes with NOTCH1 to reprogram chromatin in T-cell acutelymphoblastic leukemia
Islam R. et al.
Runt-related transcription factor 1 (RUNX1) is oncogenic in diverse types of leukemia and epithelial cancers where its expression is associated with poor prognosis. Current models suggest that RUNX1 cooperates with other oncogenic factors (e.g., NOTCH1, TAL1) to drive the expression of proto-oncogenes in T cell...Incomplete transcriptional dosage compensation of vertebrate sexchromosomes is balanced by post-transcriptional compensation
Lister N. C. et al.
Heteromorphic sex chromosomes (XY or ZW) present problems of gene dosage imbalance between the sexes, and with the autosomes. Mammalian X chromosome inactivation was long thought to imply a critical need for dosage compensation in vertebrates. However, the universal importance of sex chromosome dosage compensation w...Gene Regulatory Interactions at Lamina-Associated Domains
Madsen-Østerbye J. et al.
The nuclear lamina provides a repressive chromatin environment at the nuclear periphery. However, whereas most genes in lamina-associated domains (LADs) are inactive, over ten percent reside in local euchromatic contexts and are expressed. How these genes are regulated and whether they are able to interact with regu...Repression and 3D-restructuring resolves regulatory conflicts inevolutionarily rearranged genomes.
Ringel A. et al.
Regulatory landscapes drive complex developmental gene expression, but it remains unclear how their integrity is maintained when incorporating novel genes and functions during evolution. Here, we investigated how a placental mammal-specific gene, Zfp42, emerged in an ancient vertebrate topologically associated domai...HOTAIR interacts with PRC2 complex regulating the regional preadipocytetranscriptome and human fat distribution.
Kuo Feng-Chih et al.
Mechanisms governing regional human adipose tissue (AT) development remain undefined. Here, we show that the long non-coding RNA HOTAIR (HOX transcript antisense RNA) is exclusively expressed in gluteofemoral AT, where it is essential for adipocyte development. We find that HOTAIR interacts with polycomb repressive ...Local euchromatin enrichment in lamina-associated domains anticipatestheir repositioning in the adipogenic lineage.
Madsen-Østerbye J. et al.
BACKGROUND: Interactions of chromatin with the nuclear lamina via lamina-associated domains (LADs) confer structural stability to the genome. The dynamics of positioning of LADs during differentiation, and how LADs impinge on developmental gene expression, remains, however, elusive. RESULTS: We examined changes in t...Loss of KMT2C reprograms the epigenomic landscape in hPSCsresulting in NODAL overexpression and a failure of hemogenic endotheliumspecification.
Maurya Shailendra et al.
Germline or somatic variation in the family of KMT2 lysine methyltransferases have been associated with a variety of congenital disorders and cancers. Notably, -fusions are prevalent in 70\% of infant leukaemias but fail to phenocopy short latency leukaemogenesis in mammalian models, suggesting additional factors ar...Functional annotations of three domestic animal genomes provide vitalresources for comparative and agricultural research.
Kern C. et al.
Gene regulatory elements are central drivers of phenotypic variation and thus of critical importance towards understanding the genetics of complex traits. The Functional Annotation of Animal Genomes consortium was formed to collaboratively annotate the functional elements in animal genomes, starting with domesticate...Long intergenic non-coding RNAs regulate human lung fibroblast function: Implications for idiopathic pulmonary fibrosis.
Hadjicharalambous MR, Roux BT, Csomor E, Feghali-Bostwick CA, Murray LA, Clarke DL, Lindsay MA
Phenotypic changes in lung fibroblasts are believed to contribute to the development of Idiopathic Pulmonary Fibrosis (IPF), a progressive and fatal lung disease. Long intergenic non-coding RNAs (lincRNAs) have been identified as novel regulators of gene expression and protein activity. In non-stimulated cells, we o...Global distribution of DNA hydroxymethylation and DNA methylation in chronic lymphocytic leukemia.
Wernig-Zorc S, Yadav MP, Kopparapu PK, Bemark M, Kristjansdottir HL, Andersson PO, Kanduri C, Kanduri M
BACKGROUND: Chronic lymphocytic leukemia (CLL) has been a good model system to understand the functional role of 5-methylcytosine (5-mC) in cancer progression. More recently, an oxidized form of 5-mC, 5-hydroxymethylcytosine (5-hmC) has gained lot of attention as a regulatory epigenetic modification with prognostic ...The Itaconate Pathway Is a Central Regulatory Node Linking Innate Immune Tolerance and Trained Immunity
Domínguez-Andrés Jorge, Novakovic Boris, Li Yang, Scicluna Brendon P., Gresnigt Mark S., Arts Rob J.W., Oosting Marije, Moorlag Simone J.C.F.M., Groh Laszlo A., Zwaag Jelle, Koch Rebecca M., ter Horst Rob, Joosten Leo A.B., Wijmenga Cisca, Michelucci Ales
Sepsis involves simultaneous hyperactivation of the immune system and immune paralysis, leading to both organ dysfunction and increased susceptibility to secondary infections. Acute activation of myeloid cells induced itaconate synthesis, which subsequently mediated innate immune tolerance in human monocytes. In con...Increased H3K9 methylation and impaired expression of Protocadherins are associated with the cognitive dysfunctions of the Kleefstra syndrome.
Iacono G, Dubos A, Méziane H, Benevento M, Habibi E, Mandoli A, Riet F, Selloum M, Feil R, Zhou H, Kleefstra T, Kasri NN, van Bokhoven H, Herault Y, Stunnenberg HG
Kleefstra syndrome, a disease with intellectual disability, autism spectrum disorders and other developmental defects is caused in humans by haploinsufficiency of EHMT1. Although EHMT1 and its paralog EHMT2 were shown to be histone methyltransferases responsible for deposition of the di-methylated H3K9 (H3K9me2), th...A long range distal enhancer controls temporal fine-tuning of PAX6 expression in neuronal precursors.
Lacomme M, Medevielle F, Bourbon HM, Thierion E, Kleinjan DJ, Roussat M, Pituello F, Bel-Vialar S
Proper embryonic development relies on a tight control of spatial and temporal gene expression profiles in a highly regulated manner. One good example is the ON/OFF switching of the transcription factor PAX6 that governs important steps of neurogenesis. In the neural tube PAX6 expression is initiated in neural proge...Inhibition of Methyltransferase Setd7 Allows the In Vitro Expansion of Myogenic Stem Cells with Improved Therapeutic Potential.
Judson RN, Quarta M, Oudhoff MJ, Soliman H, Yi L, Chang CK, Loi G, Vander Werff R, Cait A, Hamer M, Blonigan J, Paine P, Doan LTN, Groppa E, He W, Su L, Zhang RH, Xu P, Eisner C, Low M, Barta I, Lewis CB, Zaph C, Karimi MM, Rando TA, Rossi FM
The development of cell therapy for repairing damaged or diseased skeletal muscle has been hindered by the inability to significantly expand immature, transplantable myogenic stem cells (MuSCs) in culture. To overcome this limitation, a deeper understanding of the mechanisms regulating the transition between ac...Metabolic Induction of Trained Immunity through the Mevalonate Pathway.
Bekkering S, Arts RJW, Novakovic B, Kourtzelis I, van der Heijden CDCC, Li Y, Popa CD, Ter Horst R, van Tuijl J, Netea-Maier RT, van de Veerdonk FL, Chavakis T, Joosten LAB, van der Meer JWM, Stunnenberg H, Riksen NP, Netea MG
Innate immune cells can develop long-term memory after stimulation by microbial products during infections or vaccinations. Here, we report that metabolic signals can induce trained immunity. Pharmacological and genetic experiments reveal that activation of the cholesterol synthesis pathway, but not the synthesis of...Krox20 hindbrain regulation incorporates multiple modes of cooperation between cis-acting elements
Thierion E. et al.
Developmental genes can harbour multiple transcriptional enhancers that act simultaneously or in succession to achieve robust and precise spatiotemporal expression. However, the mechanisms underlying cooperation between cis-acting elements are poorly documented, notably in vertebrates. The mouse gene Krox20 encodes ...A transcription factor pulse can prime chromatin for heritable transcriptional memory
Iberg-Badeaux A. et al.
Short-term and long-term transcriptional memory is the phenomenon whereby the kinetics or magnitude of gene induction is enhanced following a prior induction period. Short-term memory persists within one cell generation or in post-mitotic cells, while long-term memory can survive multiple rounds of cell division. We...β-Glucan Reverses the Epigenetic State of LPS-Induced Immunological Tolerance
Novakovic B. et al.
Innate immune memory is the phenomenon whereby innate immune cells such as monocytes or macrophages undergo functional reprogramming after exposure to microbial components such as lipopolysaccharide (LPS). We apply an integrated epigenomic approach to characterize the molecular events involved in LPS-induced to...Enhancer decommissioning by Snail1-induced competitive displacement of TCF7L2 and down-regulation of transcriptional activators results in EPHB2 silencing
Schnappauf O et al.
Transcriptional silencing is a major cause for the inactivation of tumor suppressor genes, however, the underlying mechanisms are only poorly understood. The EPHB2 gene encodes a receptor tyrosine kinase that controls epithelial cell migration and allocation in intestinal crypts. Through its ability to restrict cell...cChIP-seq: a robust small-scale method for investigation of histone modifications
Valensisi C et al.
Background ChIP-seq is highly utilized for mapping histone modifications that are informative about gene regulation and genome annotations. For example, applying ChIP-seq to histone modifications such as H3K4me1 has facilitated generating epigenomic maps of putative enhancers. This powerful technology, however, i...C/EBPα Activates Pre-existing and De Novo Macrophage Enhancers during Induced Pre-B Cell Transdifferentiation and Myelopoiesis
van Oevelen C, Collombet S, Vicent G, Hoogenkamp M, Lepoivre C, Badeaux A, Bussmann L, Sardina JL, Thieffry D, Beato M, Shi Y, Bonifer C, Graf T
Highlights C/EBPα activates two classes of prospective myeloid enhancers in B cells Pre-existing enhancers are bound by PU.1 and become hyper-activated by C/EBPα C/EBPα acts as a pioneer factor with delayed kinetics on de novo enhancers The two types of enhancers direct myeloid cell fat...SNAIL1 combines competitive displacement of ASCL2 and epigenetic mechanisms to rapidly silence the EPHB3 tumor suppressor in colorectal cancer.
Rönsch K, Jägle S, Rose K, Seidl M, Baumgartner F, Freihen V, Yousaf A, Metzger E, Lassmann S, Schüle R, Zeiser R, Michoel T, Hecht A
EPHB3 is a critical cellular guidance factor in the intestinal epithelium and an important tumor suppressor in colorectal cancer (CRC) whose expression is frequently lost at the adenoma-carcinoma transition when tumor cells become invasive. The molecular mechanisms underlying EPHB3 silencing are incompletely underst...Interrogation of allelic chromatin states in human cells by high-density ChIP-genotyping.
Light N, Adoue V, Ge B, Chen SH, Kwan T, Pastinen T
Allele-specific (AS) assessment of chromatin has the potential to elucidate specific cis-regulatory mechanisms, which are predicted to underlie the majority of the known genetic associations to complex disease. However, development of chromatin landscapes at allelic resolution has been challenging since sites of var...Nuclear ARRB1 induces pseudohypoxia and cellular metabolism reprogramming in prostate cancer
Zecchini V, Madhu B, Russell R, Pértega-Gomes N, Warren A, Gaude E, Borlido J, Stark R, Ireland-Zecchini H, Rao R, Scott H, Boren J, Massie C, Asim M, Brindle K, Griffiths J, Frezza C, Neal DE, Mills IG
Tumour cells sustain their high proliferation rate through metabolic reprogramming, whereby cellular metabolism shifts from oxidative phosphorylation to aerobic glycolysis, even under normal oxygen levels. Hypoxia-inducible factor 1A (HIF1A) is a major regulator of this process, but its activation under normoxic con...A novel microscopy-based high-throughput screening method to identify proteins that regulate global histone modification levels.
Baas R, Lelieveld D, van Teeffelen H, Lijnzaad P, Castelijns B, van Schaik FM, Vermeulen M, Egan DA, Timmers HT, de Graaf P
Posttranslational modifications of histones play an important role in the regulation of gene expression and chromatin structure in eukaryotes. The balance between chromatin factors depositing (writers) and removing (erasers) histone marks regulates the steady-state levels of chromatin modifications. Here we describe...An In-Depth Characterization of the Major Psoriasis Susceptibility Locus Identifies Candidate Susceptibility Alleles within an HLA-C Enhancer Element.
Clop A, Bertoni A, Spain SL, Simpson MA, Pullabhatla V, Tonda R, Hundhausen C, Di Meglio P, De Jong P, Hayday AC, Nestle FO, Barker JN, Bell RJ, Capon F, Trembath RC
Psoriasis is an immune-mediated skin disorder that is inherited as a complex genetic trait. Although genome-wide association scans (GWAS) have identified 36 disease susceptibility regions, more than 50% of the genetic variance can be attributed to a single Major Histocompatibility Complex (MHC) locus, known as PSORS...Disease-Related Growth Factor and Embryonic Signaling Pathways Modulate an Enhancer of TCF21 Expression at the 6q23.2 Coronary Heart Disease Locus.
Miller CL, Anderson DR, Kundu RK, Raiesdana A, Nürnberg ST, Diaz R, Cheng K, Leeper NJ, Chen CH, Chang IS, Schadt EE, Hsiung CA, Assimes TL, Quertermous T
Coronary heart disease (CHD) is the leading cause of mortality in both developed and developing countries worldwide. Genome-wide association studies (GWAS) have now identified 46 independent susceptibility loci for CHD, however, the biological and disease-relevant mechanisms for these associations remain elusive. Th...Balancing of histone H3K4 methylation states by the Kdm5c/SMCX histone demethylase modulates promoter and enhancer function.
Outchkourov NS, Muiño JM, Kaufmann K, van Ijcken WF, Groot Koerkamp MJ, van Leenen D, de Graaf P, Holstege FC, Grosveld FG, Timmers HT
The functional organization of eukaryotic genomes correlates with specific patterns of histone methylations. Regulatory regions in genomes such as enhancers and promoters differ in their extent of methylation of histone H3 at lysine-4 (H3K4), but it is largely unknown how the different methylation states are specifi...Characterization of the contradictory chromatin signatures at the 3' exons of zinc finger genes.
Blahnik KR, Dou L, Echipare L, Iyengar S, O'Geen H, Sanchez E, Zhao Y, Marra MA, Hirst M, Costello JF, Korf I, Farnham PJ
The H3K9me3 histone modification is often found at promoter regions, where it functions to repress transcription. However, we have previously shown that 3' exons of zinc finger genes (ZNFs) are marked by high levels of H3K9me3. We have now further investigated this unusual location for H3K9me3 in ZNF genes. Neither ...Using ChIP-Seq Technology to Generate High-Resolution Profiles of Histone Modifications
O’Geen H, Echipare L, Farnham PJ
The dynamic modification of DNA and histones plays a key role in transcriptional regulation through - altering the packaging of DNA and modifying the nucleosome surface. These chromatin states, also referred to as the epigenome, are distinctive for different tissues, developmental stages, and disease states and can ... - Related products


