Reagents
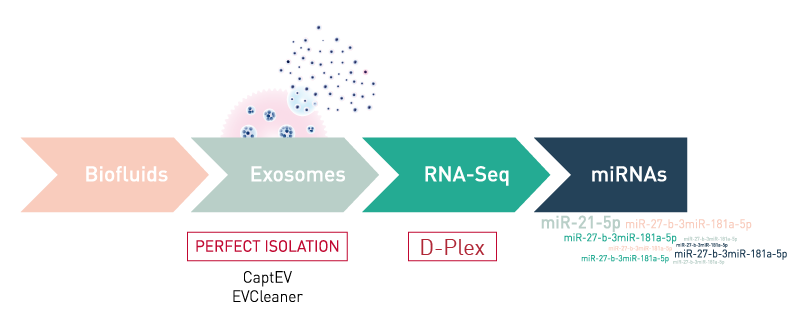
Exosomes
The identification of reliable predictive or diagnostic biomarkers is crucial for biomedical research. Recent evidence shows that exosomes, small vesicles that are excreted into the extracellular environment, are excellent disease biomarker candidates as they contain specific biological material including small non-coding RNAs (microRNAs and others) and proteins involved in intercellular communication.
Diagenode has developed specific tools for the most efficient exosome capture from biofluids (e.g. plasma, serum, urine, ...).
Benefits of our optimal exosome capture solutions
- Pure exosome capture - zero contamination from vesicles or aggregates
- Fully compatible with our D-Plex technology (D-Plex Small RNA-seq kit)
- High yields from exosome pre-enrichment
- Global and specific exosome isolation options
- Guaranteed results with high level of validation
- User-friendlly method - no ultracentrifugation needed
- Broad applications - RNA and microRNA (miRNA) NGS analysis, protein analysis, Western blot, ELISA, flow cytometry
microRNAs highlighted after exosomes isolation
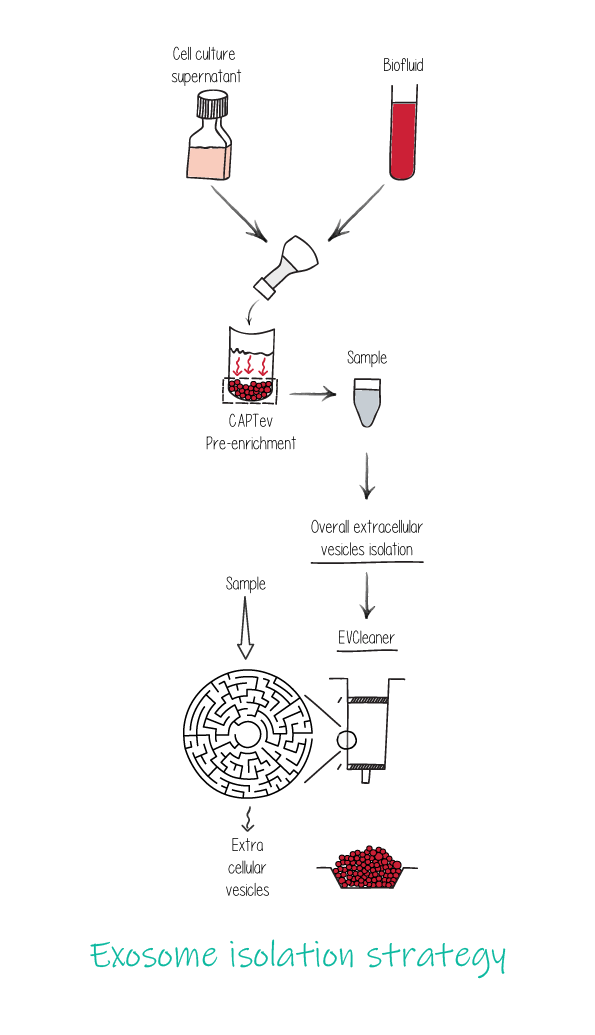
MicroRNA differential expression analysis was performed on RNA-seq data from RNA isolated from exosomes captured from HEK293T cell culture supernatant and total RNA isolated from HEK293T cells.
The RNA libraries were prepared using Diagenode's RNA-seq library preparation protocol for small RNA analysis. Reference genomes were obtained from the UCSC genome browser.
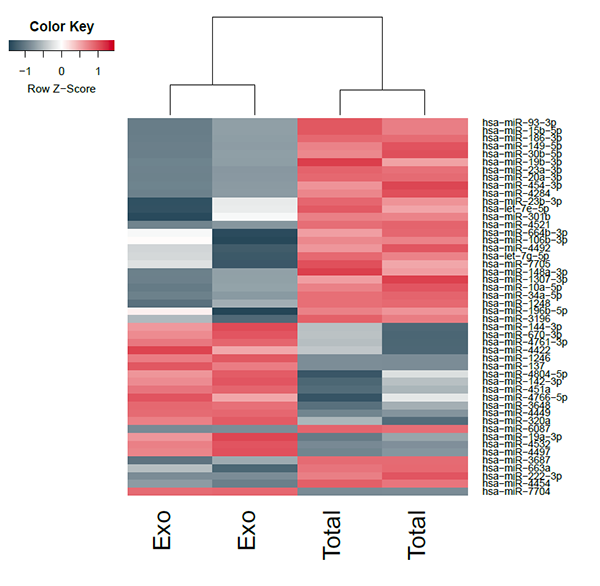
We observed several upregulated and downregulated microRNAs between exosomes isolated from cell culture supernatant and total HEK293T cells.
- microRNA upregulation including miR7704, mirR4532, miR6087, miR4497, miR3687, miR19a-3p
- microRNA downregulation including miR663a, miR4454, miR222-3p, miR196b-5p, miR10a-5p
Products
| Cat. No. | Product | Format | Price | |
|---|---|---|---|---|
| C28030001-10 |
CaptEV cell culture CaptEV is a pre-enrichement reagent that concentrates all extracellular vesicules of cell culture supernatant to ensure optimal isolation and purif... |
10 ml | $285.00 | |
| C28030002-10 |
CaptEV serum/plasma CaptEV is a pre-enrichement reagent that concentrates all extracellular vesicules of serum or plasma blood samples to ensure optimal isolation and ... |
10 ml | $285.00 | |
| C28020001-1 |
EVCleaner - DISCONTINUED EVCleaner size exclusion columns ensure efficent separation of extracellular vesicules from unwanted size components in sample. |
1 pc |