When doing CRISPR experiments, it is essential to verify the expression levels of the Cas9 to ensure high-quality results. Diagenode's new CRISPR/Cas9 4G10 antibody is extremely specific, able to recognize different Cas9 fusion proteins (even those used for CRISPRi experiments) and compatible with many biochemical assays, such as immunofluorescence, western blot and immunoprecipitation. Among the different antibodies in the market, Diagenode's 4G10 CRISPR/Cas9 is our antibody of choice.
Researcher at EPFL, Lausanne, Switzerland
CRISPR/Cas9 Antibody 4G10 (sample size)
Catalog Number
Format
Price
Other format
| Lot | 004 |
|---|---|
| Concentration | 1.4 μg/μl |
| Species reactivity | Streptococcus pyogenes |
| Type | Monoclonal |
| Purity | Protein G purified monoclonal antibody in PBS containing 0.05 % Na-azide. |
| Host | Mouse |
| Storage Conditions | Store at -20°C; for long storage, store at -80°C. Avoid multiple freeze-thaw cycles. |
| Precautions | This product is for research use only. Not for use in diagnostic or therapeutic procedures. |
| Applications | Suggested dilution * | References |
|---|---|---|
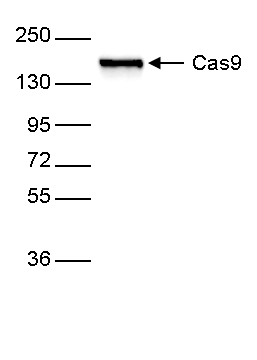
| Western Blotting | 1:5,000 | Fig 1 |
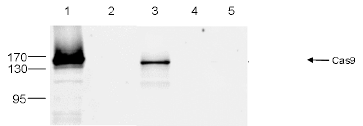
| Immunoprecipitation | 5 μg/IP | Fig 2 |
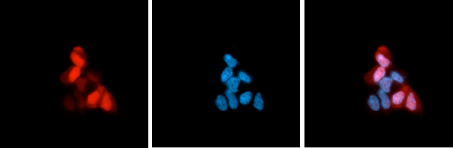
| Immunofluorescence | 1:400 | Fig 3 |