D-Plex mRNA-seq Kit for Illumina

(Coming soon)
D-Plex mRNA-seq Library Preparation Kit is a tool designed for the study of the whole coding transcriptome. The kit is using the D-Plex technology to generate directionnal libraries for Illumina sequencing directly from total RNAs or mRNAs that has already been enriched by poly(A) selection.
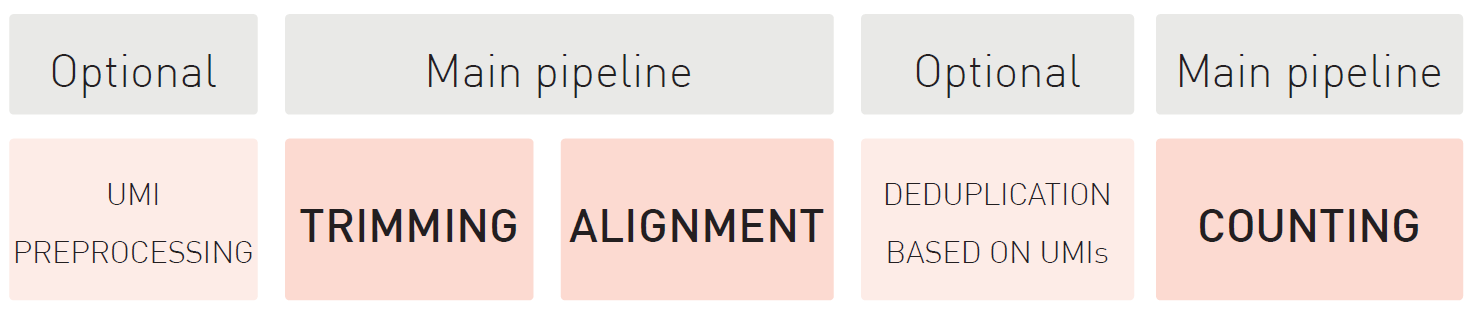
The D-Plex technology utilizes the innovative capture and amplification by tailing and switching, a ligation-free method for RNA library preparation from ultra-low input amounts. Combined with unique molecular identifiers (UMI), this complete solution delivers a high-sensitivity detection method for low-abundance or rare transcripts.
D-Plex mRNA-seq Kit offers a time saving protocol that can be completed within 5 hours and requires minimal hands-on time. The library preparation takes place in a single tube, increasing the efficiency tremendously. This new solution has been extensively validated for both intact and highly degraded RNA samples, including that derived from FFPE preparations.
D-Plex Total RNA-seq Kit includes all buffers and enzymes necessary for the library preparation. Specific D-Plex Unique Dual Indexes were designed and validated to fit the D-Plex technology and are available separately: