This antibody specifically recognizes the AML1 (RUNX1) (UniProtKB/Swiss-Prot entry Q01196) - ETO (RUNX1T1) (UniProtKB/ Swiss-Prot entry Q06455) fusion protein that arises due to a translocation between chromosome 8 and 22 (t(8;21)(q22;q22)). This translocation is one of the most frequent karyotypic abnormalities observed in acute myeloid leukaemia. It produces a chimerical gene made up of the 5’-region of AML1and the 3’-region of ETO. The chimerical protein is thought to associate with the nuclear corepressor/histone deacetylase complex to block hematopoietic differentiation.
AML1-ETO Antibody - ChIP-seq Grade (sample size)
| Lot | A706-002 |
|---|---|
| Concentration | not determined |
| Species reactivity | Human |
| Type | Polyclonal |
| Purity | Whole antiserum |
| Host | Rabbit |
| Precautions | This product is for research use only. Not for use in diagnostic or therapeutic procedures. |
| Applications | Suggested dilution | References |
|---|---|---|
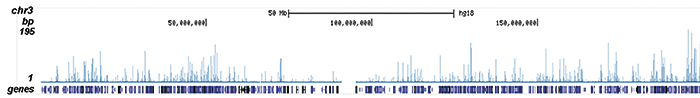
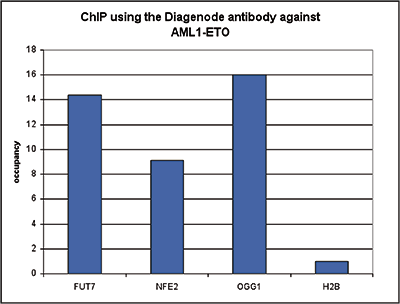
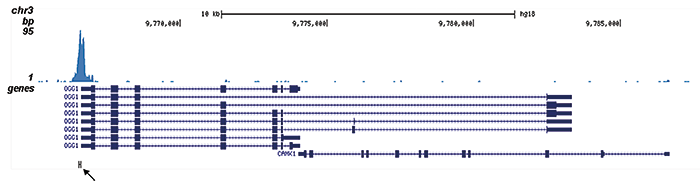
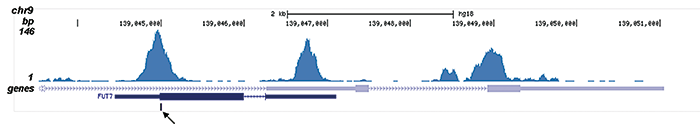
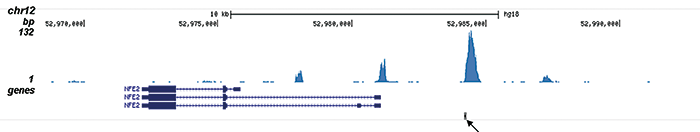
| ChIP/ChIP-seq * | 4 μl/ChIP | Fig 1, 2 |
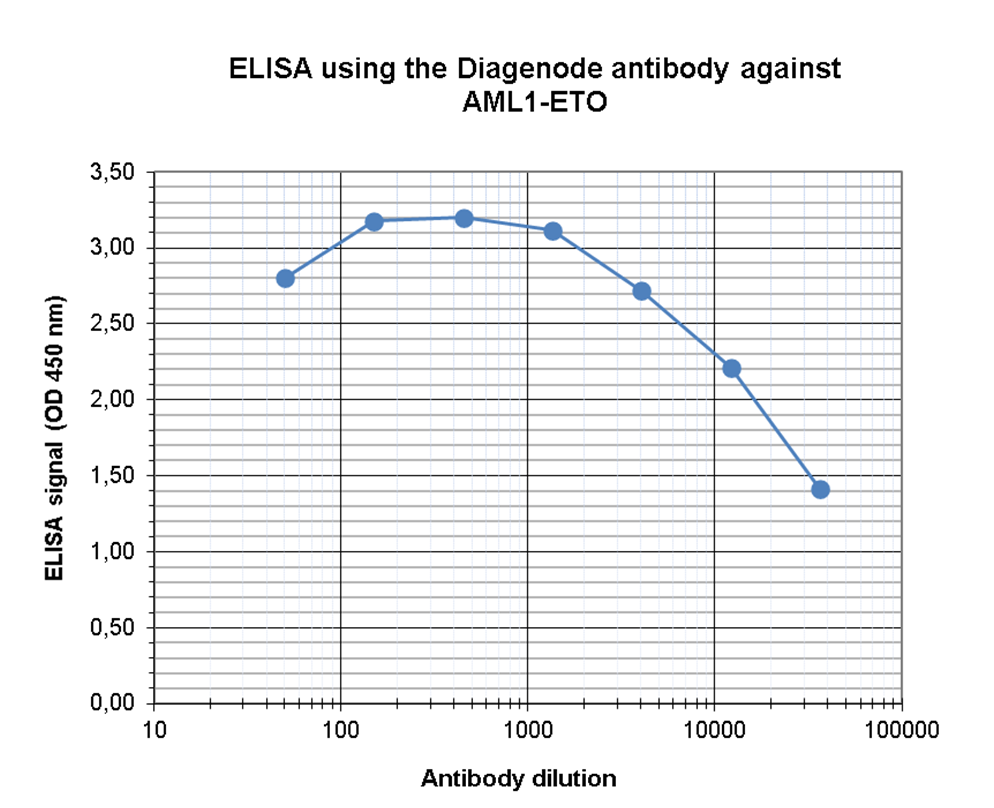
| ELISA | 1:500 | Fig 3 |
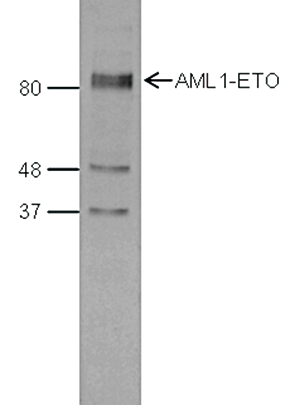
| Western Blotting | 1:1,000 | Fig 4 |
* Please note that the optimal antibody amount per ChIP should be determined by the end-user. We recommend testing 1-10 μl per IP.