| Automating Whole-Genome DNA Methylation Analyses POSTER A step toward identifying subtle epigenetic variation in cancer and other complex disease | Download |
| MagMeDIP qPCR Kit - User Manual v2 MANUAL This manual concerns: MagMeDIP x48 batch #15 and before; MagMeDIP x10 batch #7 a... | Download |
| MagMeDIP qPCR Kit - User Manual v4 MANUAL This manual concerns: MagMeDIP x48 batch #16 and after; MagMeDIP x10 batch #7A and af... | Download |
Auto MagMeDIP qPCR Kit
Perform MeDIP (Methylated DNA Immunoprecipitation) on the SX-8G IP-Star® Automated System followed by qPCR to estimate DNA methylation status of your sample using 5-methylcytosine antibody. Our kit contains high quality reagents to get the highest enrichment of methylated DNA with an optimized user-friendly protocol.
Diagenode’s Auto MagMeDIP qPCR is available in two formats (10 and 48 IPs) and has been optimized on the SX-8G IP-Star® Automated System enabling highly reproducible results and allowing for high throughput assays.
Characteristics
- Generate highly consistent results with internal controls in 24h
- Minimize error with many reagents in 1 tube
- Optimized purification (DIB - DNA isolation buffer)
- Allows direct correlation between IP’d material & methylation status
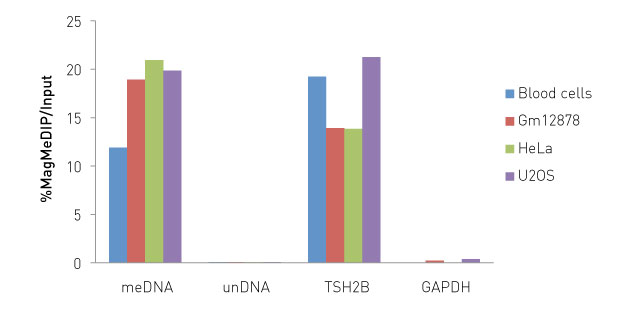
Figure 1. IP results obtained with Diagenode Auto MagMeDIP qPCR Kit. MeDIP assays were performed manually using DNA from blood, Gm12878, Hela and U20S cells and the Auto MagMeDIP qPCR kit (Diagenode). The DNA was prepared with the XL GenDNA Extraction Module included. The IP was performed including the kit meDNA and unDNA spike-in controls, together with the human DNA sample controls. The DNA was isolated/purified using DIB. Afterwards, qPCR was performed using the primer pairs also included in this kit.
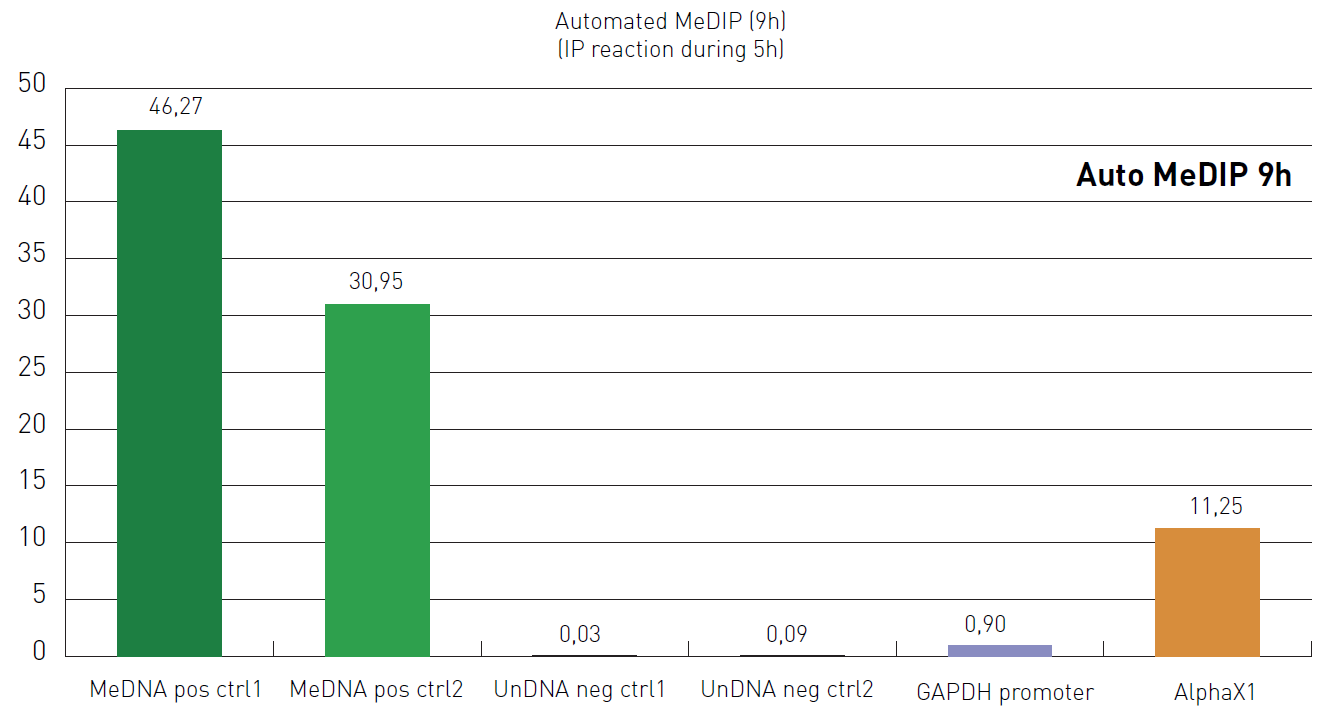
Figure 2. Automated MeDIP (9h). IP reaction was performed on the SX-8G IP-Star® Automated System with the anti-5-mC antibody. Methylated and unmethylated DNA were used as internal controls. Unmethylated DNA region of GADPH and a methylated DNA region of AlphaX1 were used to test DNA sample-IP efficiency. DNA has been isolated by using DNA Isolation Buffer (DIB).
