DESCRIPTION
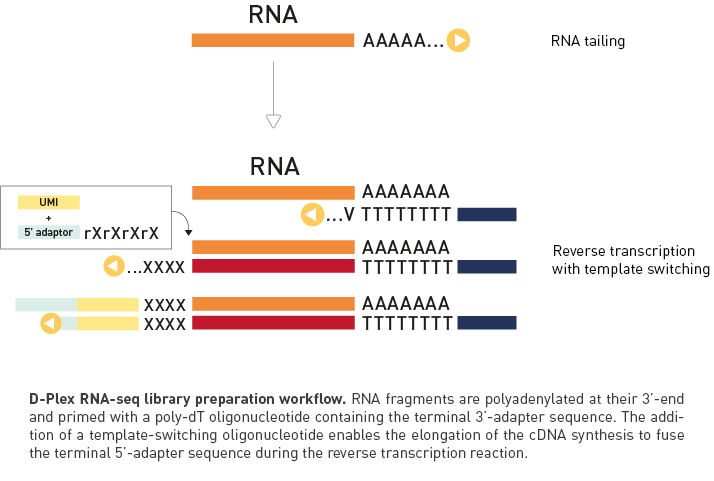
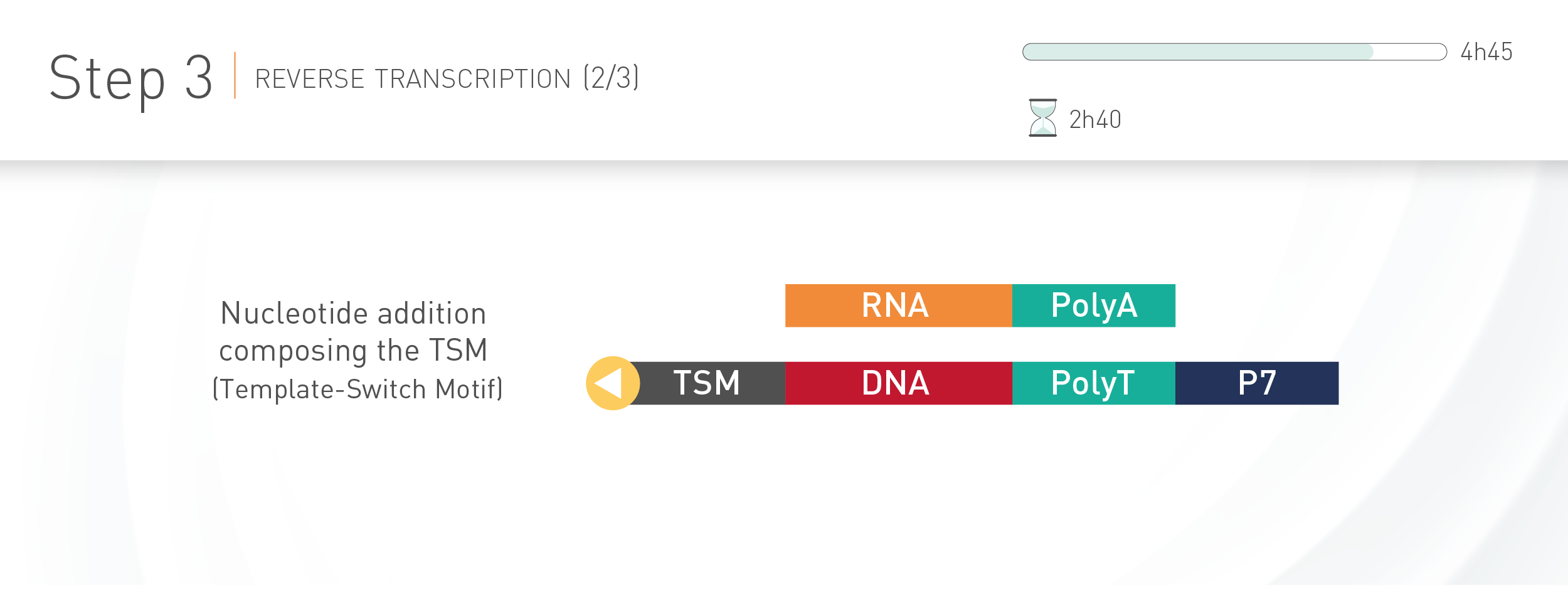
D-Plex technology combines RNA tailing and template-switching
The D-Plex technology utilizes the innovative capture and amplification by tailing and switching, a unique, ultrasensitive ligation-free method to generate RNA libraries for next generation sequencing. This ingenious enzymatic technology offers key advantages compared to ligase-based approach:
- Low to ultra-low input capability with high reproducibility
- Higher library complexity providing novel and diverse transcript detection
- Optimal performance on challenging samples, such as liquid biopsies and FFPE samples
The D-Plex technology offers new RNA-seq solution, optimized and specifically formatted for both Illumina’s and MGI’s sequencing platforms.
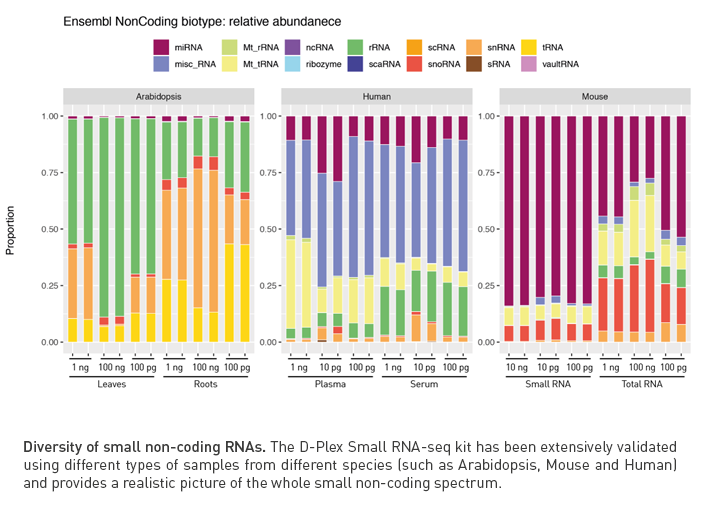
D-Plex provides high transcript diversity
Thanks to the template-switching mechanism, D-Plex RNA-seq solution ensures a realistic representation of the transcriptome of your sample of interest.
The D-Plex Small RNA-seq kit and the D-Plex Small RNA DNBSEQ kit efficiently incorporate a wide variety of small RNA species, such as microRNAs (miRNAs), PIWI-interacting RNAs (piRNAs), small nucleolar RNAs (snoRNAs), small nuclear RNAs (snRNAs) and tRNA-derived small RNAs, which represent new potential classes of biomarkers to be explored as they have proved to be key components for cellular regulation in some diseases.
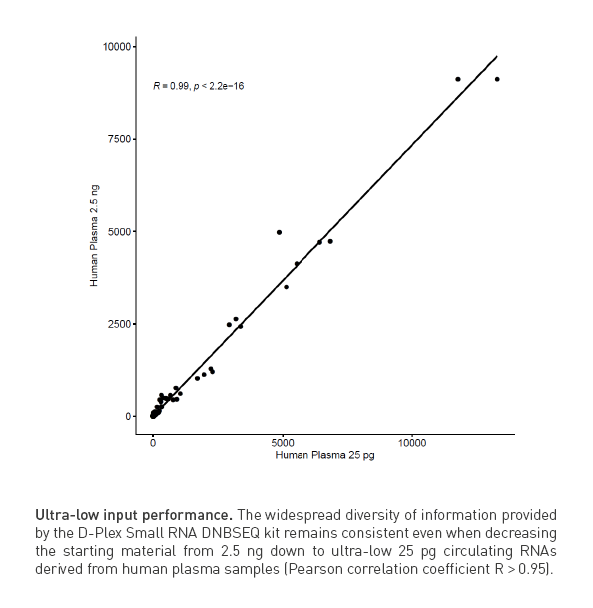
D-Plex is highly suitable for low RNA inputs and challenging samples
The input requirements for D-Plex solutions are flexible and allow the user to perform the library preparation within a wide range of RNA quantities going from 10 pg of small RNAs (< 200 nt) or circulating RNAs up to 100 ng of total RNAs.
The D-Plex RNA-seq kits have been extensively validated for clinically relevant samples such as liquid biopsies or FFPE samples. The D-Plex technology is exceedingly suitable for epigenetic biomarker discovery.
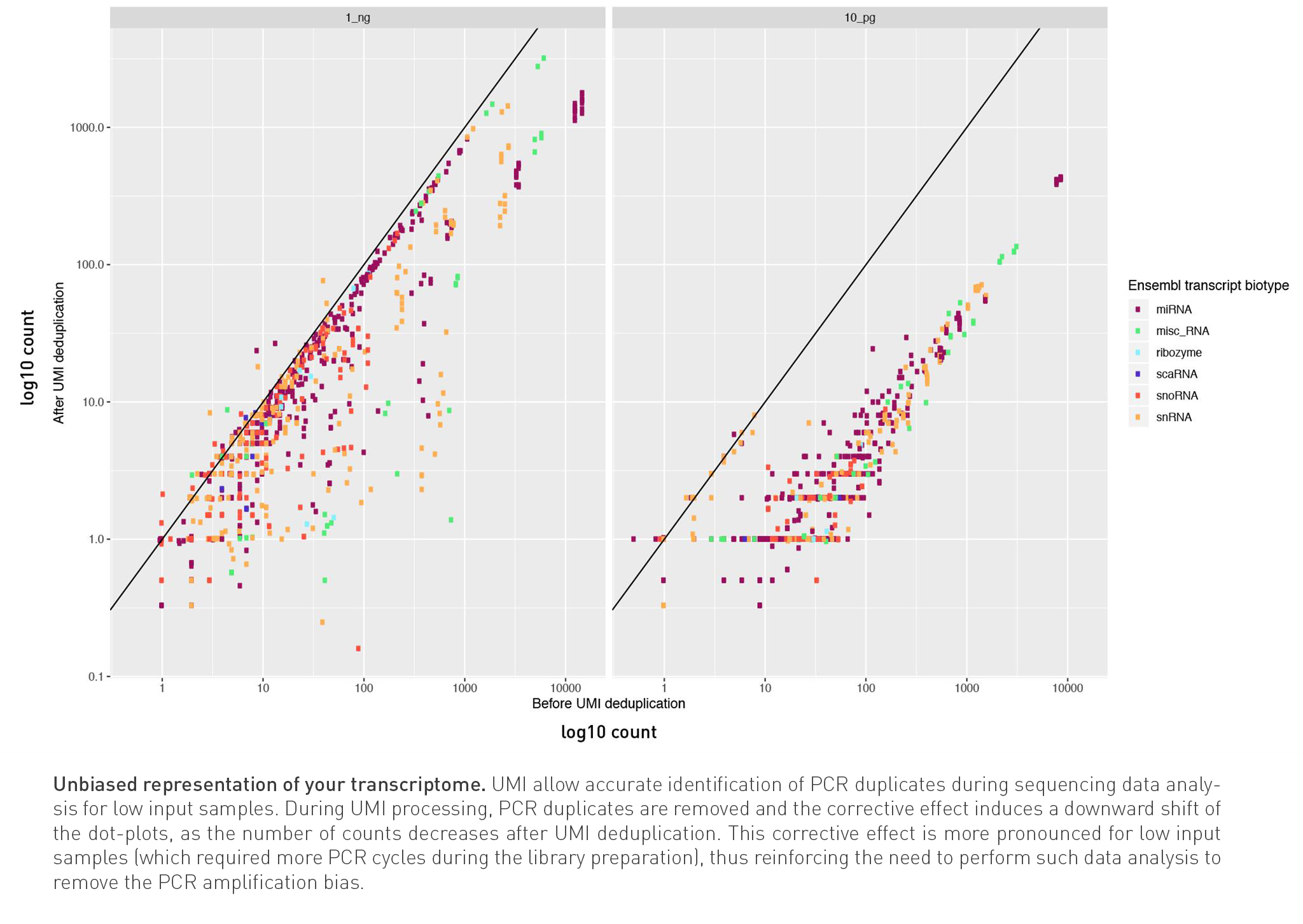
UMI allow accurate identification of duplicates for low input samples
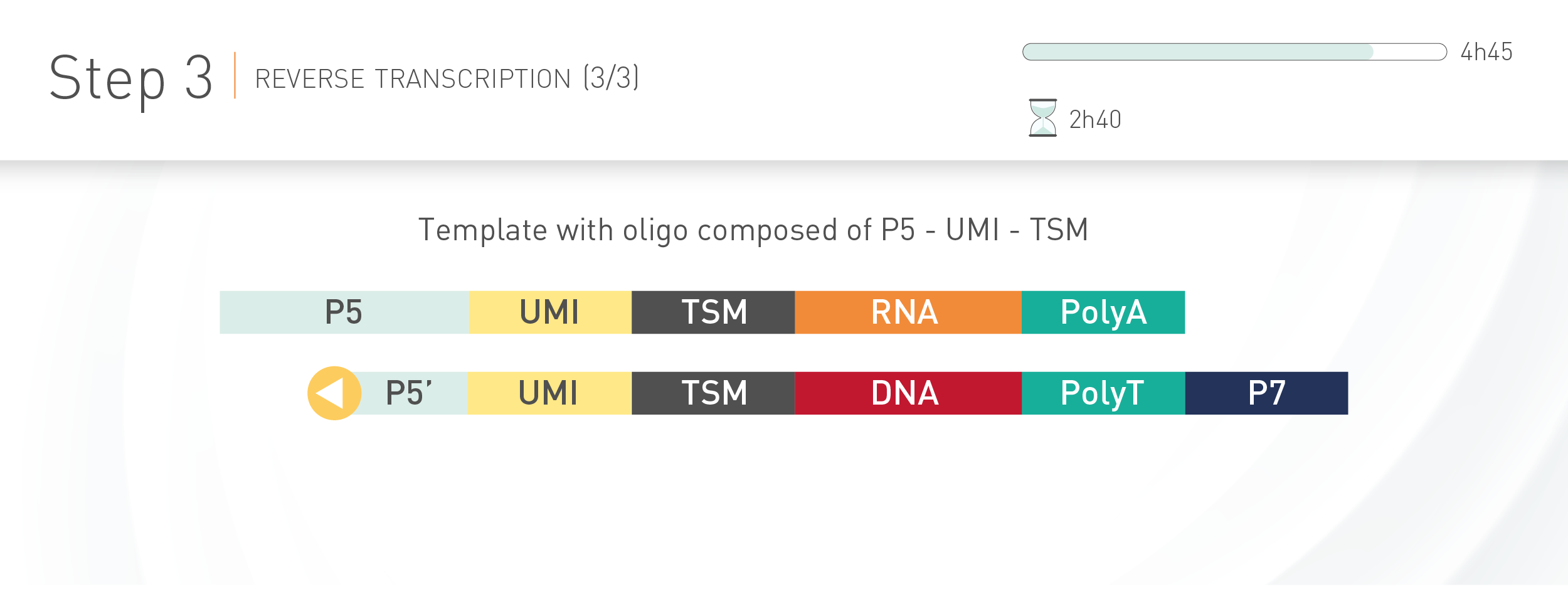
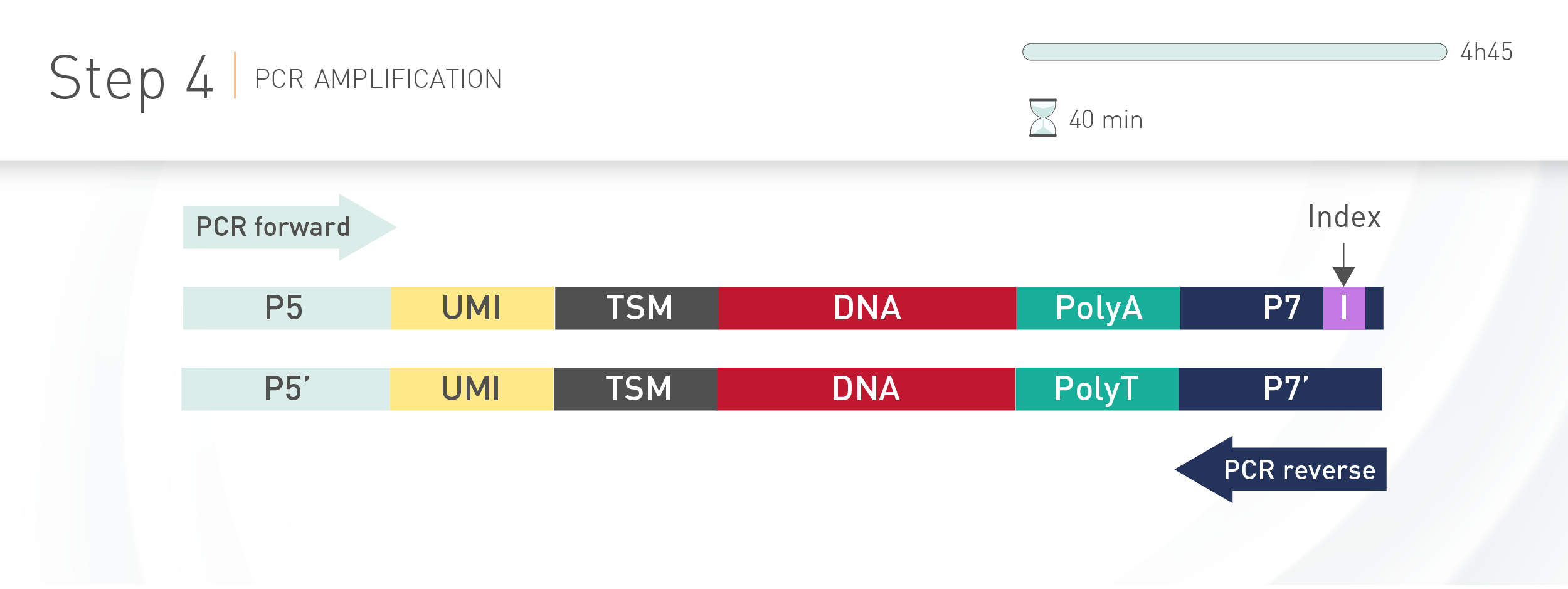
D-Plex technology for Illumina sequencing integrates unique molecular identifiers (UMI) to each transcript incorporated in the final library. The use of such molecular tag gives the possibility to exclude PCR duplicates from a set of reads during sequencing data analysis, thus improving the transcript expression quantification.
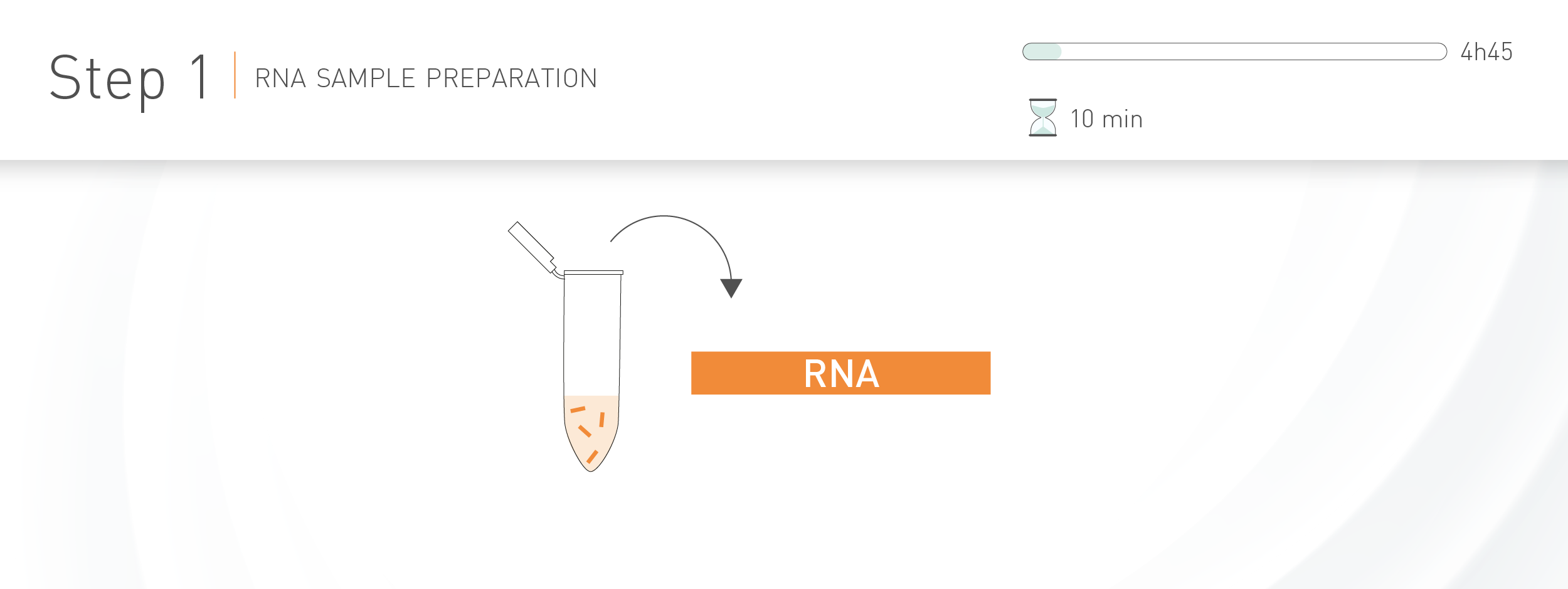
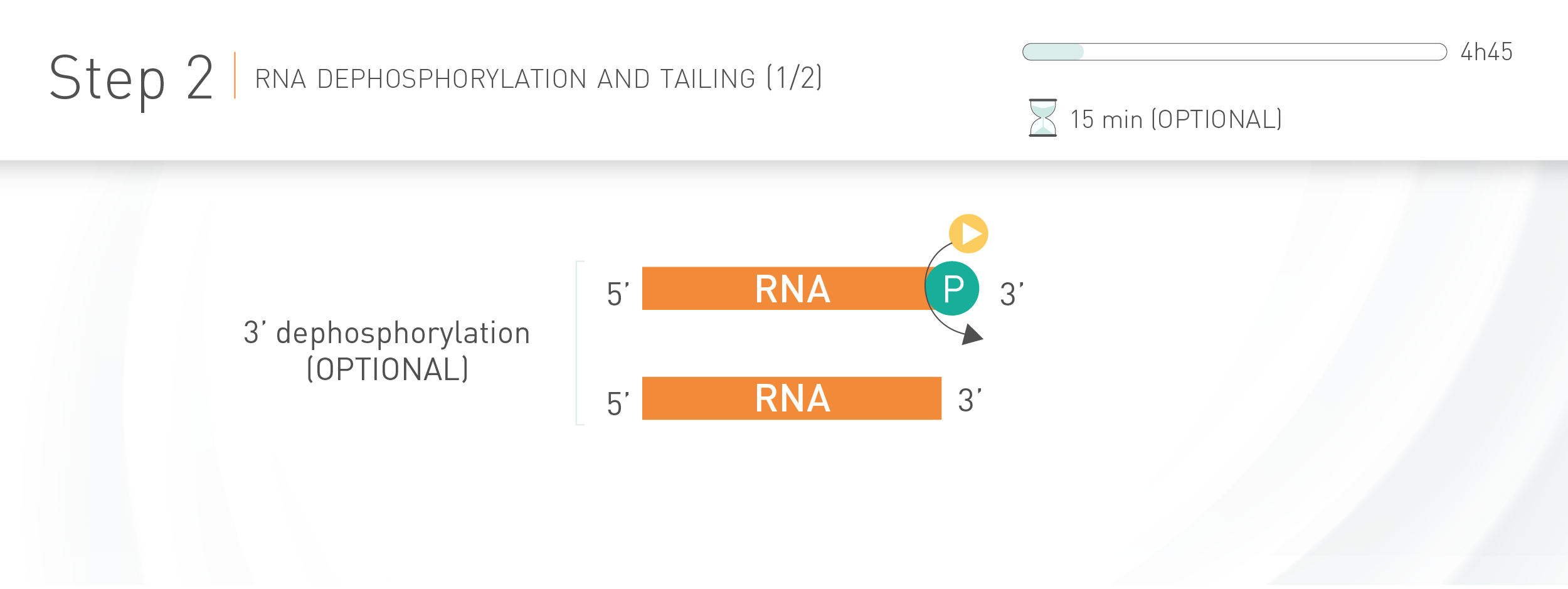
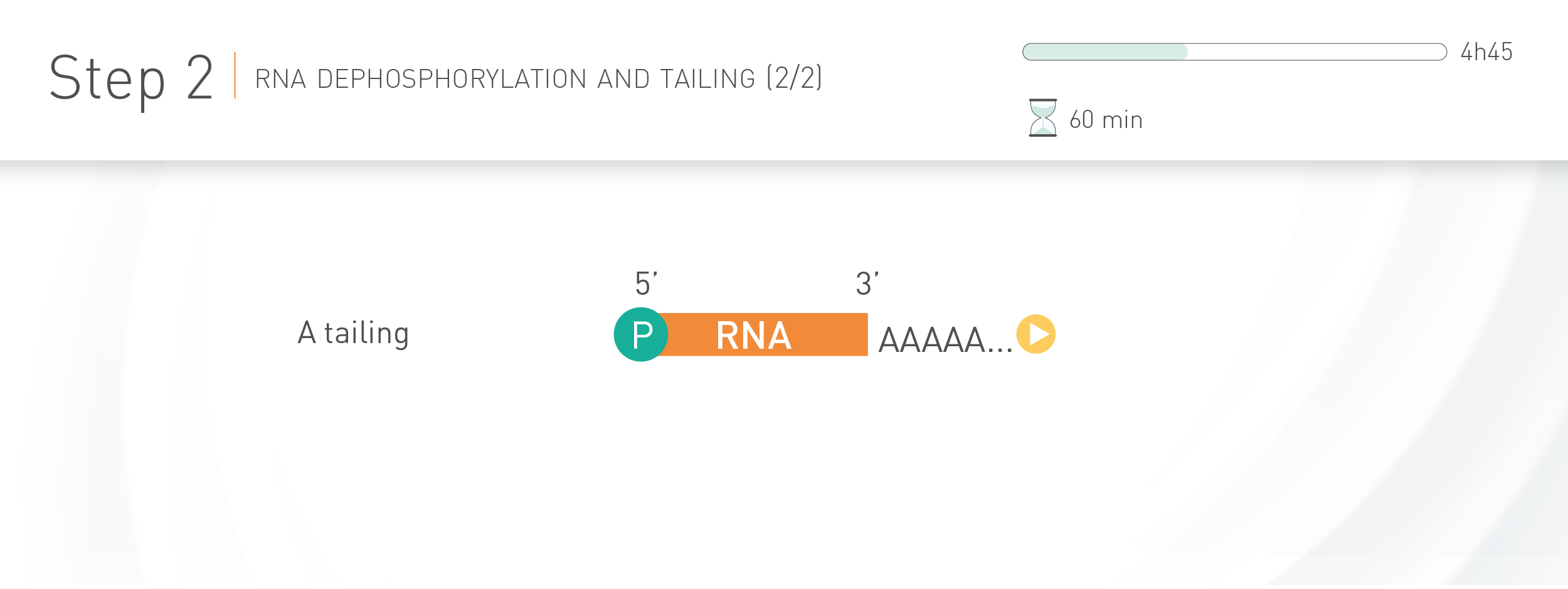
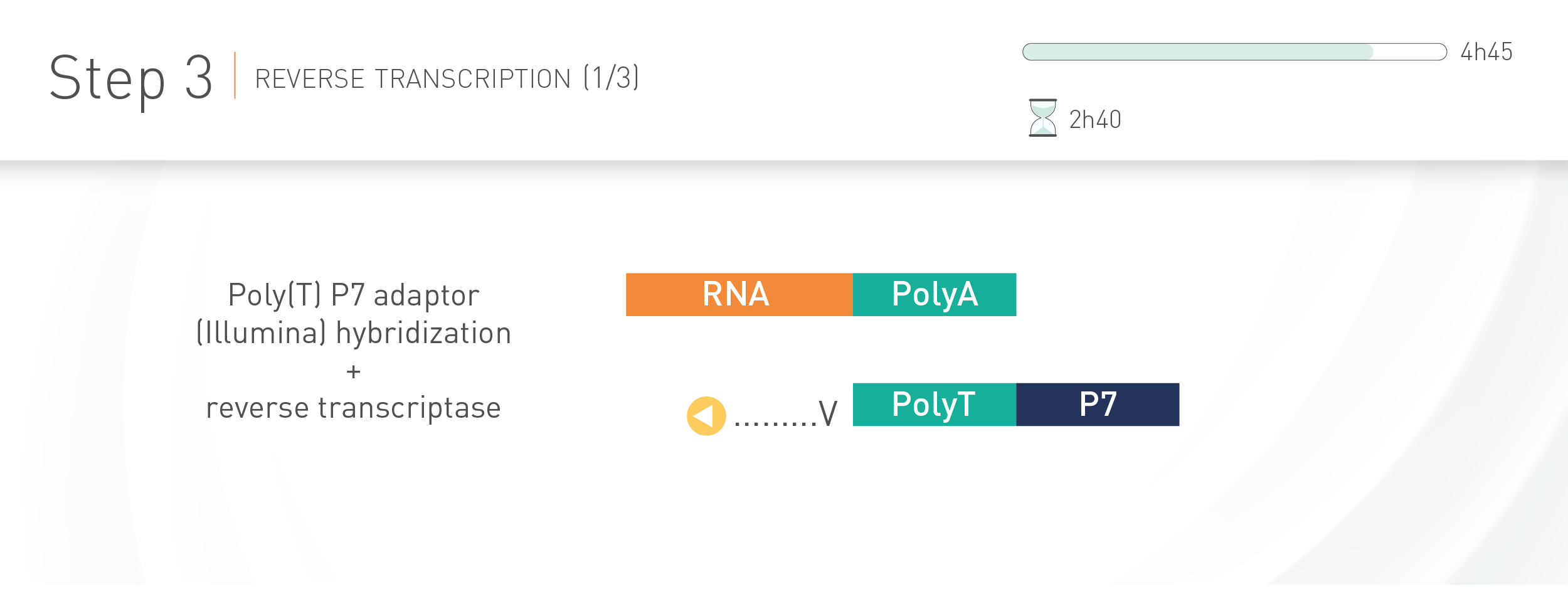
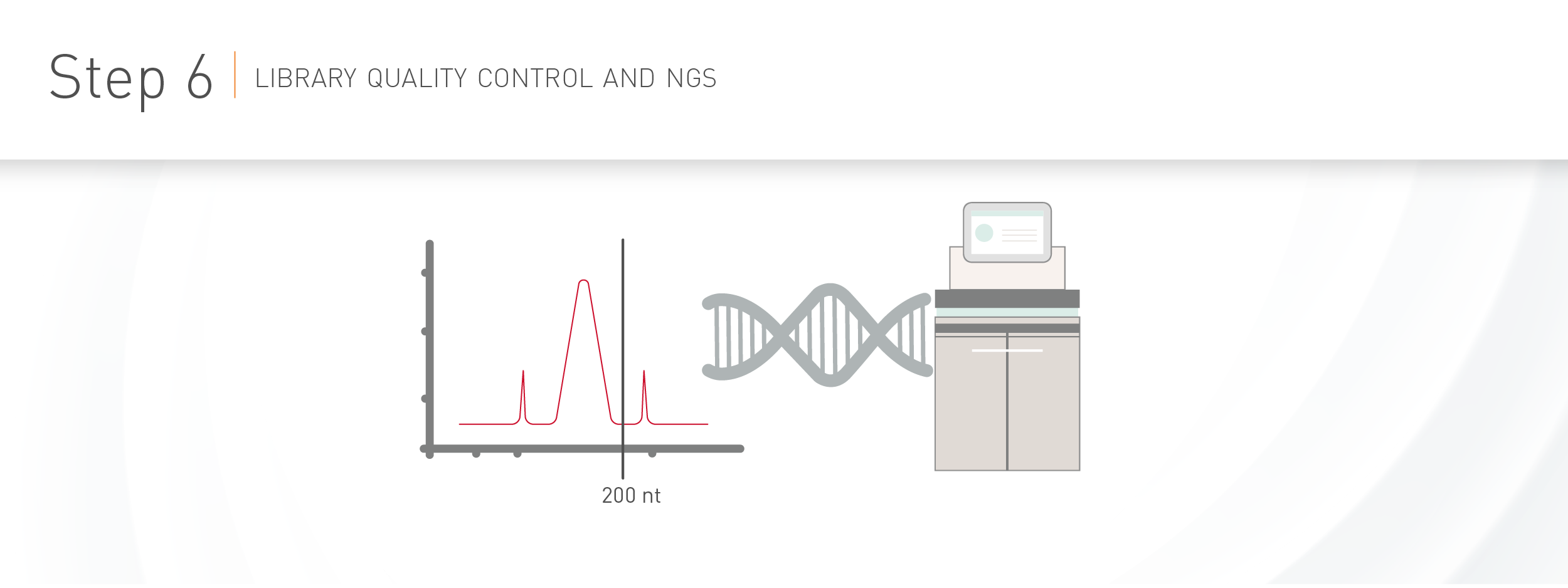
WORKFLOW
SOLUTIONS
TESTIMONIAL
They trust us !
“We work with extremely limited quantities of small RNAs. In our hands, D-Plex had the highest yield compared to many other approaches. Inclusion of UMIs is essential for our small RNA analyses.”
Can Cenik, PhD